Naxerova Lab
Read BioOur lab’s research spans a variety of topics in cancer genetics and somatic evolution. We use interdisciplinary approaches to study the origins and consequences of somatic variation. We blend computational analysis, experimental work and mathematical modeling - whatever it takes!
We are interested in understanding how processes of mutation and selection in normal (stem) cells set the stage for cancer evolution over long periods of time. Then, once a tumor develops, how does tissue-specific selection shape the cancer genome? How can we take advantage of genetic intra-tumor heterogeneity to gain insights into the life history of a cancer? Finally, the evolution of metastasis is a particular focus of the lab. Do metastases arise from distinct clones with special, genetically encoded properties or do they represent random samples of the primary tumor? Does metastatic spread happen early or late in tumor development? Do all metastases arise independently from the primary tumor, or do they give rise to each other? How heterogeneous are metastases?
The answers to the questions above have important clinical implications but are difficult to study in human patients because it is challenging to reconstruct occult events that happened years before diagnosis. We have developed genetic techniques to determine the clonal architecture and lineage of cancer cells in human specimens and collaborate with clinicians in utilizing these tools to further our understanding of cancer evolution.
Research topics:
- Experimental investigation and mathematical modeling of somatic evolution in normal tissues and cancers
- Cancer phylogenetics
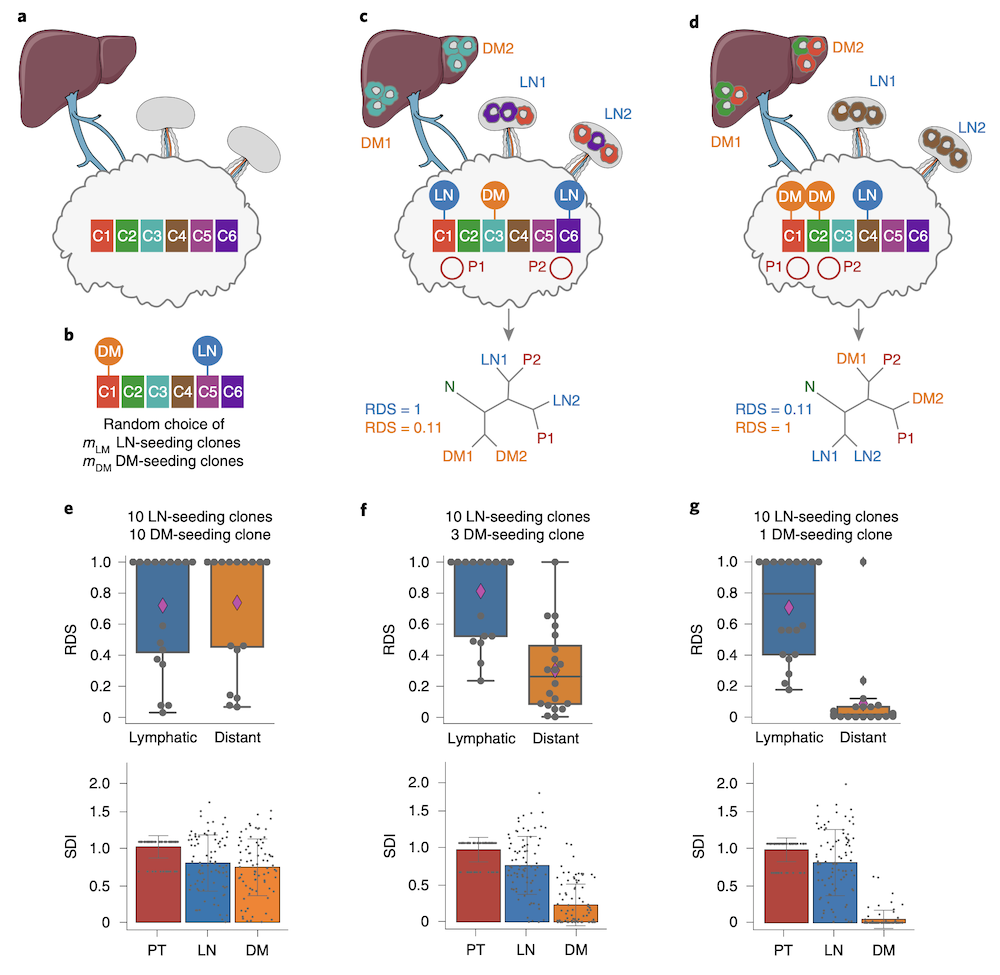
- Metastasis evolution
- High-throughput genetic screening
- Integrative data analysis
Recent Publications
Zhang S, Paccalet A, Rohde D, Cremer S, Hulsmans M, Lee IH, Mentkowski K, Grune J, Schloss MJ, Honold L, Iwamoto Y, Zheng Y, Bredella MA, Buckless C, Ghoshhajra B, Thondapu V, van der Laan AM, Piek JJ, Niessen HWM, Pallante F, Carnevale R, Perrotta S, Carnevale D, Iborra-Egea O, Muñoz-Guijosa C, Galvez-Monton C, Bayes-Genis A, Vidoudez C, Trauger SA, Scadden D, Swirski FK, Moskowitz MA, Naxerova K, Nahrendorf M Bone marrow adipocytes fuel emergency hematopoiesis after myocardial infarction. Nat Cardiovasc Res. 2023;2(12):1277-1290 - PMID: 38344689 - PMCID: PMC10857823 - DOI: 10.1038/s44161-023-00388-7
Hulsmans M, Schloss MJ, Lee IH, Bapat A, Iwamoto Y, Vinegoni C, Paccalet A, Yamazoe M, Grune J, Pabel S, Momin N, Seung H, Kumowski N, Pulous FE, Keller D, Bening C, Green U, Lennerz JK, Mitchell RN, Lewis A, Casadei B, Iborra-Egea O, Bayes-Genis A, Sossalla S, Ong CS, Pierson RN, Aster JC, Rohde D, Wojtkiewicz GR, Weissleder R, Swirski FK, Tellides G, Tolis G, Melnitchouk S, Milan DJ, Ellinor PT, Naxerova K, Nahrendorf M Recruited macrophages elicit atrial fibrillation. Science. 2023;381(6654):231-239 - PMID: 37440641 - PMCID: PMC10448807 - DOI: 10.1126/science.abq3061
Janizek JD, Dincer AB, Celik S, Chen H, Chen W, Naxerova K, Lee SI Uncovering expression signatures of synergistic drug responses via ensembles of explainable machine-learning models. Nat Biomed Eng. 2023;7(6):811-829 - PMID: 37127711 - PMCID: PMC11149694 - DOI: 10.1038/s41551-023-01034-0
Küçükköse E, Laoukili J, Gorelick AN, Degner S, Lacle M, van den Bent L, Peters NA, Verheem A, Hung WT, Frenkel NC, Wassenaar E, Lansu N, Lenos KJ, Vermeulen L, Koopman M, Roodhart JML, Kops GJPL, Borel Rinkes IHM, Hagendoorn J, Naxerova K, Kranenburg O Lymphatic invasion of plakoglobin-dependent tumor cell clusters drives formation of polyclonal lung metastases in colon cancer. Gastroenterology. 2023;165(2):429-444.e15 - PMID: 36906044 - DOI: 10.1053/j.gastro.2023.02.047
Datta M, Chatterjee S, Perez EM, Gritsch S, Roberge S, Duquette M, Chen IX, Naxerova K, Kumar AS, Ghosh M, Emblem KE, Ng MR, Ho WW, Kumar P, Krishnan S, Dong X, Speranza MC, Neagu MR, Iorgulescu JB, Huang RY, Youssef G, Reardon DA, Sharpe AH, Freeman GJ, Suvà ML, Xu L, Jain RK Losartan controls immune checkpoint blocker-induced edema and improves survival in glioblastoma mouse models. Proc Natl Acad Sci U S A. 2023;120(6):e2219199120 - PMID: 36724255 - PMCID: PMC9963691 - DOI: 10.1073/pnas.2219199120
- More publications ...
News
Kamila Naxerova received the Emerging Leader Award from the The Mark Foundation for Cancer Research. The Emerging Leader Award program empowers scientists to take on innovative, high-risk/high-reward projects that have significant potential to improve outcomes for cancer patients. Congratulations, Kamila!
Kamila Naxerova has been awarded the AACR NextGen Grants for Transformative Cancer Research. This award represents the AACR’s flagship funding initiative to stimulate highly innovative research from young investigators. Congratulations, Kamila!
The Naxerova lab is looking for two new team members to study metastasis patterns in colorectal cancer and melanoma. Please read more details about the available positions here.