Lee Lab
Read BioOur vision is to create cutting-edge platform technologies that detect disease at their earliest stages through better analysis of biological systems. We leverage our expertise in nanomaterials, biophysics, electronics, and computation to achieve this goal, creating robust engineering solutions. We have made significant progress in developing and refining new biosensors optimized for specific medical and research needs. Many of these sensors have been tested in clinical trials and are translated for commercialization.

Recent Publications
Woo HK, Kim C, Choi Y, Cho YK, Do LN, Kim H, Jung DH, Allen M, Jeon J, Chung S, Park SY, Park I, Castro CM, Park JS, Lee H
Automated disc device for multiplexed extracellular vesicle isolation and labelling from liquid biopsies in cancer diagnostics. Nat Biomed Eng. 2026;10(5):882-896 - PMID: 41606293 - DOI: 10.1038/s41551-025-01601-7Di Ianni E, Jeon J, Mohammadi S, Hu H, Quintana JM, Lee C, Haidar EA, Goemans M, Zargani-Piccardi A, Mahamdeh M, Hernández IC, Rodriguez VE, Zhou Y, Aguirre A, Yuan S, Ng TSC, Breyne K, Lee H, Breakefield XO, Miller MA
Enhanced-mRNA Delivery Using Ultrasound-Delivered Anchors for Bioorthogonal Ligation. Angew Chem Int Ed Engl. 2026;65(10):e23437 - PMID: 41568620 - PMCID: PMC13052994 - DOI: 10.1002/anie.202523437Park JH, Lee DH, Park EK, Cho YK, Shin IS, Lee H
Smartphone-integrated lateral flow assay for robust detection of Δ9-Tetrahydrocannabinol. Biosens Bioelectron. 2025;296:118349 - PMID: 41478036 - PMCID: PMC12950521 - DOI: 10.1016/j.bios.2025.118349Kim YI, Lee C, Lee H, Park IJ
Extracellular vesicles in colorectal cancer. Ann Coloproctol. 2025;41(5):379-392 - PMID: 41158051 - PMCID: PMC12569783 - DOI: 10.3393/ac.2025.00745.0106Jeon J, Liu QJ, Woo H, Barth I, Choi Y, Sang LJ, Lee H
Computationally Designed Nanobinders as Affinity Ligands in Diagnostic and Therapeutic Applications. J Am Chem Soc. 2025;147(35):32187-32198 - PMID: 40839867 - PMCID: PMC13181443 - DOI: 10.1021/jacs.5c11289- More publications ...
Research projects
Liquid biopsy of extracellular vesicles (EVs)
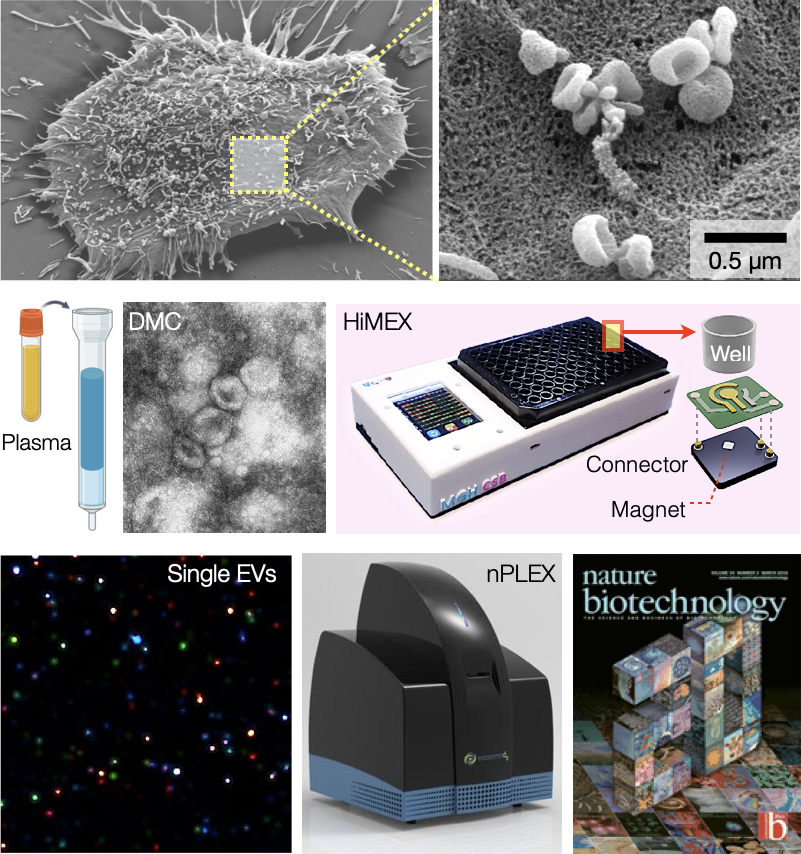 We are conducting extensive research into analyzing extracellular vesicles (EVs), membrane-bound nanoparticles released by cells. We have extensive expertise in developing tools tailored for EV analyses (please see our review article in Chemical Reviews). These include a dual-column device to enrich EVs from abundant lipoproteins, microfluidics for EV RNA analysis, and new electronic or optical sensors (HiMEX, iPEX, nPLEX) for EV protein screening. Notably, our flagship technology, nPLEX, was selected as one of the greatest hits in 20 years of Nature Biotechnology biomedical research. We are also developing an exploratory platform for multi-dimensional single EV profiling. Applying these new technologies, we have demonstrated EVs' clinical usefulness in various aspects of cancer care. Investigating EVs specific to organs can help us better understand the progression of diseases, which has far-reaching implications for detecting early cancer and relapse. Furthermore, EV mRNA testing can predict how patients will respond to treatment. Multi-institutional clinical trials are underway to validate these findings further. Ultimately, we envision establishing periodic EV tests to gain dynamic molecular information on tumors, paving the way for personalized care.
We are conducting extensive research into analyzing extracellular vesicles (EVs), membrane-bound nanoparticles released by cells. We have extensive expertise in developing tools tailored for EV analyses (please see our review article in Chemical Reviews). These include a dual-column device to enrich EVs from abundant lipoproteins, microfluidics for EV RNA analysis, and new electronic or optical sensors (HiMEX, iPEX, nPLEX) for EV protein screening. Notably, our flagship technology, nPLEX, was selected as one of the greatest hits in 20 years of Nature Biotechnology biomedical research. We are also developing an exploratory platform for multi-dimensional single EV profiling. Applying these new technologies, we have demonstrated EVs' clinical usefulness in various aspects of cancer care. Investigating EVs specific to organs can help us better understand the progression of diseases, which has far-reaching implications for detecting early cancer and relapse. Furthermore, EV mRNA testing can predict how patients will respond to treatment. Multi-institutional clinical trials are underway to validate these findings further. Ultimately, we envision establishing periodic EV tests to gain dynamic molecular information on tumors, paving the way for personalized care.
Global cancer challenge
 Cancer is an urgent global health concern with a hefty burden on low-income countries (LMICs). Two-thirds of the world's cancer deaths occur in LMICs, and the incidence rate is rapidly increasing. However, regular cancer screening and early diagnosis remain challenging due to limited medical infrastructure and high testing costs. This situation leads to unnecessary suffering and deaths, especially from breast and cervical cancer, which are easily curable if detected early. To address these pressing challenges, we have been developing fast and affordable diagnostic tools, leveraging our expertise in system engineering. Our portfolio includes an AI-powered diffraction analysis (AIDA), an ultrafast plasmonic PCR system, and a CRISPR-based digital test. We are also actively engaged in the clinical application of these technologies. A cancer diagnosis trial is planned in Ghana and Uganda, supported by the Affordable Cancer Technologies Program (NIH). Through these efforts, we are committed to reducing regional disparities in medical diagnosis and improving the quality of life.
Cancer is an urgent global health concern with a hefty burden on low-income countries (LMICs). Two-thirds of the world's cancer deaths occur in LMICs, and the incidence rate is rapidly increasing. However, regular cancer screening and early diagnosis remain challenging due to limited medical infrastructure and high testing costs. This situation leads to unnecessary suffering and deaths, especially from breast and cervical cancer, which are easily curable if detected early. To address these pressing challenges, we have been developing fast and affordable diagnostic tools, leveraging our expertise in system engineering. Our portfolio includes an AI-powered diffraction analysis (AIDA), an ultrafast plasmonic PCR system, and a CRISPR-based digital test. We are also actively engaged in the clinical application of these technologies. A cancer diagnosis trial is planned in Ghana and Uganda, supported by the Affordable Cancer Technologies Program (NIH). Through these efforts, we are committed to reducing regional disparities in medical diagnosis and improving the quality of life.
Computational imaging and machine learning
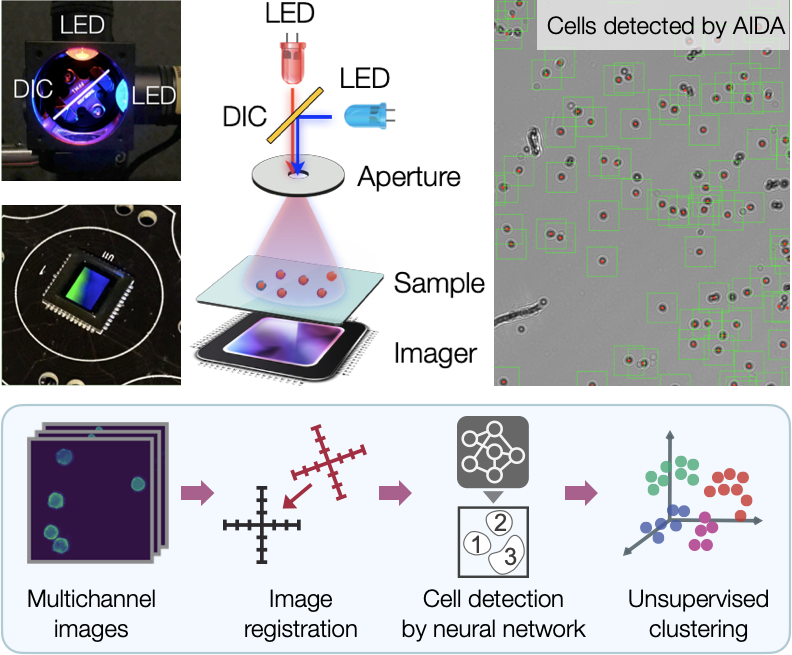 Advances in digital sensors and computing have ushered in a new age of computational optics. This technology can overcome fundamental limits in conventional optics by acquiring optically-coded images that can be numerically analyzed and transformed. We aim to develop an intelligent imaging system integrating computational optics and state-of-the-art machine vision technologies. This synergistic approach could segment and count thousands of single cells per acquisition, automatically analyze large data sets, and extract critical features (visual biomarkers). Our AIDA (AI-powered diffraction analysis) system is the first prototype, combining digital holography with deep learning for automated whole-image analyses. Our next system will be based on giga-pixel microscopy, which records a sequence of low-resolution images and then digitally restores high-resolution details. The system will create “Google Earth”-like tissue images – one can readily visualize large areas of samples and fluidly zoom up to reveal all the fine details of single cells. To ensure efficient data analysis, we have teamed up with Dr. Kwonmoo Lee, an expert in machine learning and computer vision.
Advances in digital sensors and computing have ushered in a new age of computational optics. This technology can overcome fundamental limits in conventional optics by acquiring optically-coded images that can be numerically analyzed and transformed. We aim to develop an intelligent imaging system integrating computational optics and state-of-the-art machine vision technologies. This synergistic approach could segment and count thousands of single cells per acquisition, automatically analyze large data sets, and extract critical features (visual biomarkers). Our AIDA (AI-powered diffraction analysis) system is the first prototype, combining digital holography with deep learning for automated whole-image analyses. Our next system will be based on giga-pixel microscopy, which records a sequence of low-resolution images and then digitally restores high-resolution details. The system will create “Google Earth”-like tissue images – one can readily visualize large areas of samples and fluidly zoom up to reveal all the fine details of single cells. To ensure efficient data analysis, we have teamed up with Dr. Kwonmoo Lee, an expert in machine learning and computer vision.
New membrane sensor platform
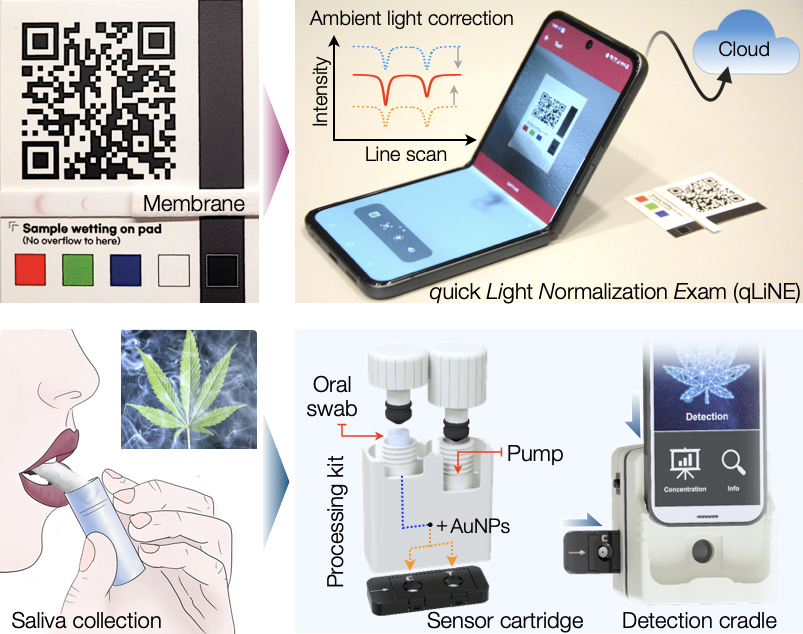 Membrane sensors are fast, cost-effective, and easy to use with minimal sample preparation, leading to their wide adoption in rapid assay kits. We are further building this platform to provide robust and qualitative molecular analyses. Our aim is to make membrane sensors as sensitive as ELISA but with a much quicker turnaround time. The qLiNE (quick Light Normalization Exam) method is a good example. It allows a regular smartphone to be used as a reliable signal reader. This system can automatically correct shape distortion, illumination brightness, and color imbalances, providing consistent optical signals in varying lighting conditions. We also developed a radial membrane technology that significantly shortens the assay time while maintaining analytical sensitivity. This method was integrated with a compact optical setup for on-site marijuana testing. Our sensor quantified tetrahydrocannabinol (THC), marijuana’s primary psychoactive substance, at the level below the regulatory guideline within 5 minutes. In a pilot field testing, we could identify recent (<12 hours) marijuana users by detecting THC in saliva samples.
Membrane sensors are fast, cost-effective, and easy to use with minimal sample preparation, leading to their wide adoption in rapid assay kits. We are further building this platform to provide robust and qualitative molecular analyses. Our aim is to make membrane sensors as sensitive as ELISA but with a much quicker turnaround time. The qLiNE (quick Light Normalization Exam) method is a good example. It allows a regular smartphone to be used as a reliable signal reader. This system can automatically correct shape distortion, illumination brightness, and color imbalances, providing consistent optical signals in varying lighting conditions. We also developed a radial membrane technology that significantly shortens the assay time while maintaining analytical sensitivity. This method was integrated with a compact optical setup for on-site marijuana testing. Our sensor quantified tetrahydrocannabinol (THC), marijuana’s primary psychoactive substance, at the level below the regulatory guideline within 5 minutes. In a pilot field testing, we could identify recent (<12 hours) marijuana users by detecting THC in saliva samples.
Emerging new technologies
 We are pioneering new signaling technologies to empower biosensors. For immunosensing, we leverage electrochemiluminescence (ECL), whose unique signaling mechanism confers competitive advantages, including exceptional sensitivity with a little optical background, consistent analytical signal through the spatiotemporal control of chemical reactions, and simpler measurement setups than other assay methods. For instance, our ECLipse (ECL in paired signal electrode) technology is highly modular and scalable, with a sensitivity up to 7000-fold higher than ELISA. In parallel, we are actively researching new strategies for nucleic acid detection. Our latest systems combine CRISPR machinery and nucleic acid amplification in a single-step reaction, significantly streamlining the assay processes while reducing the total assay time (30 min). We have further incorporated the method into a hydrogel microsystem, building a CLAMP (Cas-Loaded Annotated Micro-Particles) that allows for high-throughput and multi-target detection.
We are pioneering new signaling technologies to empower biosensors. For immunosensing, we leverage electrochemiluminescence (ECL), whose unique signaling mechanism confers competitive advantages, including exceptional sensitivity with a little optical background, consistent analytical signal through the spatiotemporal control of chemical reactions, and simpler measurement setups than other assay methods. For instance, our ECLipse (ECL in paired signal electrode) technology is highly modular and scalable, with a sensitivity up to 7000-fold higher than ELISA. In parallel, we are actively researching new strategies for nucleic acid detection. Our latest systems combine CRISPR machinery and nucleic acid amplification in a single-step reaction, significantly streamlining the assay processes while reducing the total assay time (30 min). We have further incorporated the method into a hydrogel microsystem, building a CLAMP (Cas-Loaded Annotated Micro-Particles) that allows for high-throughput and multi-target detection.
News
Fantastic news! Dr. Ju-ro Lee (Lee Lab) has been appointed to the faculty at the Department of Chemical Engineering, Dankook University, Korea. Big congratulations to Ju-ro on embarking on this new career!
Phenomenal news! Ms. Hayoung Cho (Lee Lab) has been named the sole recipient of the prestigious 2025 Edward H. Finnegan, S.J., Award as the graduating senior from Boston College. This is the highest undergraduate Commencement honor of the College. Many congratulations to Hayoung on this well-deserved recognition!
Congratulations to Dr. Jueun Jeon (Lee lab) for winning the MGH Celebration of Science 2025 Poster of Distinction Award! This marks the Lee group's third consecutive year receiving this honor.
Great start to 2025! Dr. Ala Jo, a former member of the Lee lab, has been appointed to a faculty at Ehwa Women's University. Congratulations, Ala!
Dr. Youngkwan Cho (Lee lab) has been selected as a winner of the MGH Celebration of Science 2024 Poster of Distinction Award. Many congratulations!