Cracking bacterial code for quick diagnosis
Bacterial infections remain one of the biggest killers of humans and animals. This is mostly due to the fact that it can take days to accurately identify infecting pathogens. By this point, an infection can worsen if not treated with the appropriate antibiotics to begin with. A new test developed by researchers at the CSB, however, may soon make it possible to accurately identify infecting strains in under 2 hours. This test, which uses bacterial-specific genetic material for identification, would enable clinicians to detect infections early and administer targeted antibiotics during that window of opportunity when they are still treatable.
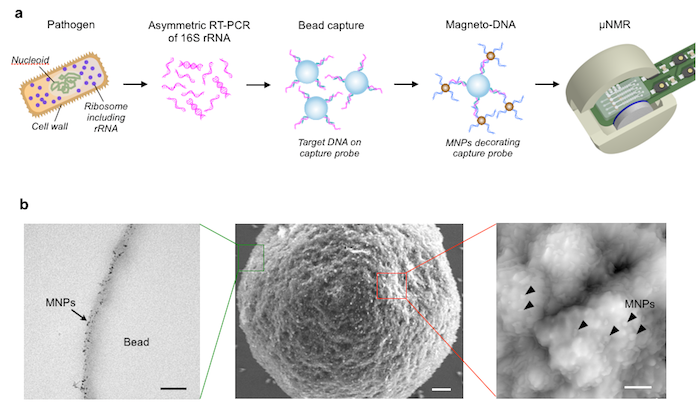
The diagnostic device, reported in the May issue of Nature Nanotechnology, uses specially designed DNA probes that seek out, capture, and magnetically label either general or species-specific regions of a particular kind of bacterial RNA known as 16S ribosomal RNA (16S rRNA). 16S rRNA is a known bacterial biomarker, and by using DNA probes targeted for either universal regions (present in all bacteria) or species-specific regions of 16S rRNA, the diagnostic test could identify both the general presence of bacteria as well as specific bacterial species responsible for infection. These DNA probes not only captured but also magnetically tagged these RNA regions, which could then be easily detected by magnetic resonance.
Magnetic resonance had been previously demonstrated at the CSB to be an excellent and accurate method for detecting magnetically-labeled biological targets. In its original design, the method employed antibodies for detection and labeling. “The problem with using antibodies for detecting bacteria was we couldn’t find any good ligands that detected the bacteria specifically,” explains Hyun Jung Chung, PhD, a post-doc at the CSB and first author on the paper. “So we thought why don’t we use a genetic detection method because that way we can design probes according to what we want to detect.”
Using this system on a series of bacterial cultures, the investigators first demonstrated that they could detect all bacteria universally as well as individual bacteria specifically, even when bacteria were combined in one sample. Once shown feasible in bacterial cultures, the investigators then tested the detection approach on clinical samples. “We were pretty excited about the potential for this technology,” confesses Cesar Castro, MD, an investigator at the CSB and co-author on the paper. “However, many promising approaches fail when translated into the clinical reality. We sought out to determine if we could leverage the benefits we saw at the benchside using real human specimens from the bedside.”
Clinical specimens were thus collected from patients with different bacterial infections, and results using the magnetic-DNA detection method were compared with gold standard techniques.
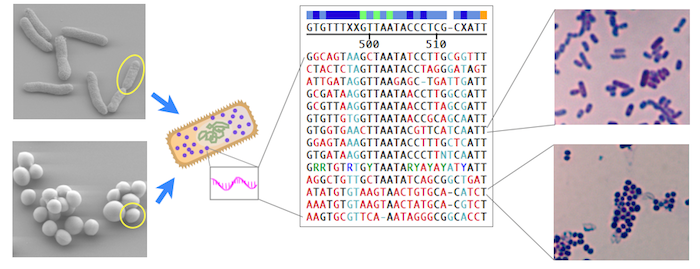
“We found that our magnetic-DNA approach in this pilot study showed excellent sensitivity and specificity, to the same level as conventional culture,” confirms Castro. The new method could even detect single bacteria within a couple of hours, compared to 3 to 5 days with conventional culture.
While these results are indeed exciting, Hakho Lee, Assistant Professor at MGH and a senior author on the paper, is quick to point out that for now, this is still a proof-of-concept. “For clinical application, we need to package everything into a stand-alone machine—contained and sealed—to minimize contamination.”
Ultimately, according to Castro, “our goal is to move this from clinical proof-of-concept to actual clinical validation. That is, to move beyond correlations with gold standards to actual clinical endpoints and establish if earlier detection with our approach confers tangible morbidity and mortality benefits. On paper, this seems very plausible.”
Written by Yvonna Fisher-Jeffes, PhD
Chung HJ, Castro CM, Im H, Lee H, Weissleder R
A magneto-DNA nanoparticle system for rapid detection and phenotyping of bacteria
Nat Nanotechnol. 2013;8(5):369-75 – PMID: 23644570 – PMCID: PMC3711657
Press Coverage
The Scientist – The Sooner, The Better (pdf)